How do we detect differences between a set of reads from a microorganism and a reference genome?
Variant calling is the process of identifying differences between two genome samples. Usually differences are limited to single nucleotide polymorphisms (SNPs) and small insertions and deletions (indels).
Imagine that you have been asked to find the differences between a sample that has been sequenced and a known genome.
For example:
You have a new sample from a patient and you want to see if it has any differences from a well known reference genome of the same species.
Typically, you would have a couple of fastq read files sent back to you from the sequencing provider and either an annotated or non annotated reference genome.
In this tutorial, we will use the tool Snippy to find high confidence differences (indels or SNPs) between our known genome and our reads, ie. finds SNPs between a haploid reference genome and your NGS sequence reads.
Snippy pipeline consists of 3 steps:
BWA MEM with a custom set of settings which are very suitable to aligning reads for microbial type dataFreebayes with a custom set of settingsSnpEff with a custom set of settingsThe Galaxy wrapper for Snippy has the ability to change some of the underlying tool settings in the advanced section but it is not recommended.
The data is a subset of a real dataset from a Staphylococcus aureus bacteria. We have a closed genome sequence and an annotation for our wildtype strain.
A whole-genome shotgun approach was used to produce a series of short sequence reads on an Illumina DNA sequencing instrument for the mutant strain.
The reads are paired-end
Each read is on average 150 bases
The reads would cover the original wildtype genome to a depth of 19x
The files we will be using are:
mutant_R1.fastq & mutant_R2.fastq - the read files in fastq format.
wildtype.fna - The sequence of the reference strain in fasta format.
wildtype.gbk - The reference strain with gene and other annotations in genbank format.
wildtype.gff - The reference strain with gene and other annotations in gff3 format.
NOTE! This data is available at Zenodo
Create a new history
Upload datasets
https://zenodo.org/record/582600/files/mutant_R1.fastq
https://zenodo.org/record/582600/files/mutant_R2.fastq
https://zenodo.org/record/582600/files/wildtype.fna
https://zenodo.org/record/582600/files/wildtype.gbk
https://zenodo.org/record/582600/files/wildtype.gff
We will now run the Snippy tool on our reads, comparing it to the reference.
Snippy is a tool for rapid bacterial SNP calling and core genome alignments.
Snippy finds SNPs between a haploid reference genome and your NGS sequence reads. It will find both substitutions (snps) and insertions/deletions (indels).
If we give Snippy an annotated reference, it will silently run SnpEff which will figure out the effect of any changes on the genes and other features.
If we just give Snippy the reference sequence alone without the annotations, it will not run SnpEff.
We have an annotated reference and so will use it in this case.
Snippy tool with the following parameters:
Snippy has taken the reads and:
mapped them against the reference using BWA MEM,
looked through the resulting BAM file and found differences using some fancy Bayesian statistics (Freebayes),
filtered the differences for sensibility and finally checked what effect these differences will have on the predicted genes and other features in the genome.
It produces quite a bit of output, there can be up to 10 output files.

snps vcf file: The final annotated variants in VCF format
snps gff file: The variants in GFF3 format
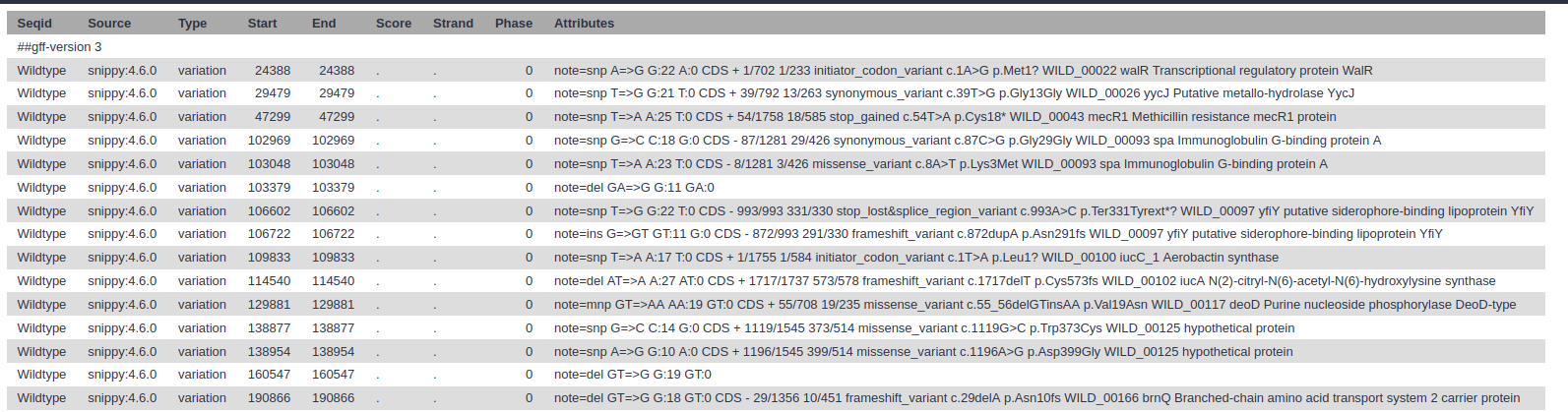
snps table: A simple tab-separated summary of all the variants
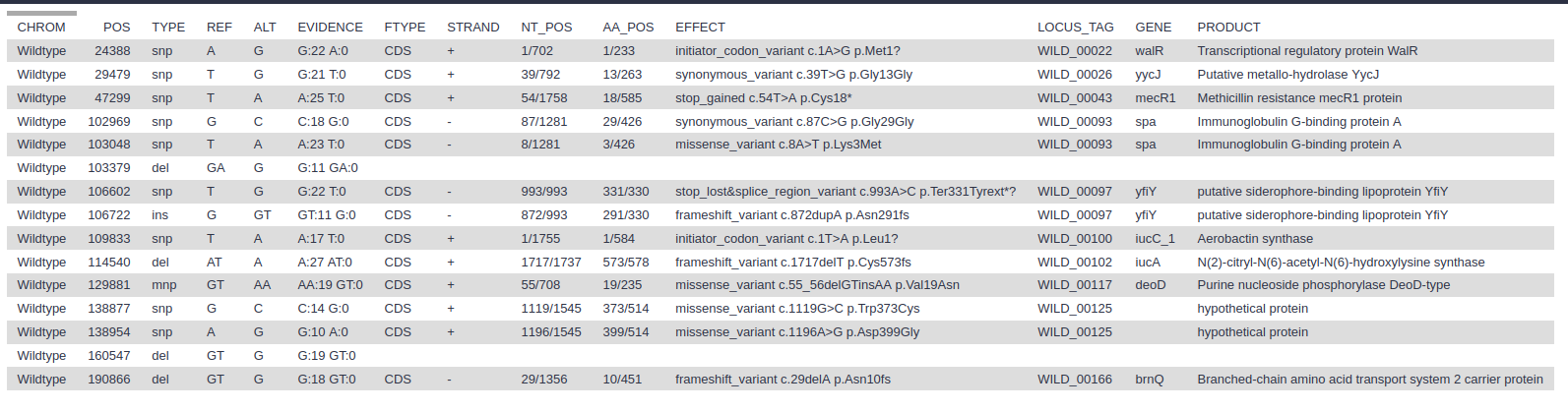
snps summary: A summary of the SNPs called
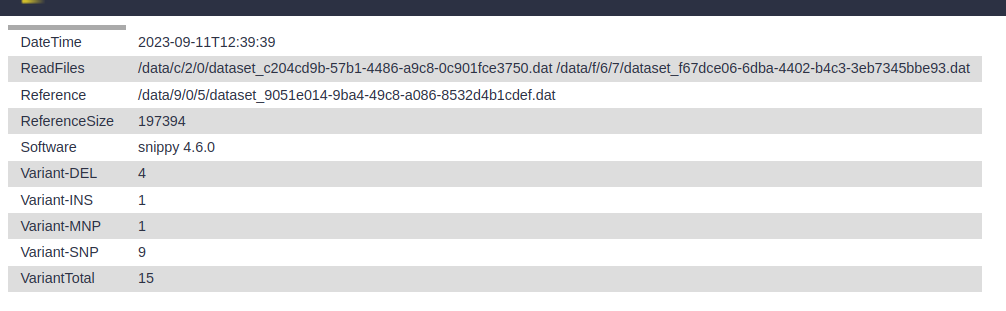
log file: A log file with the commands run and their outputs
aligned fasta: A version of the reference but with - at position with depth=0 and N for 0 < depth < –mincov (does not have variants)
consensus fasta: A version of the reference genome with all variants instantiated
mapping depth: A table of the mapping depth
mapped reads bam: A BAM file containing all of the mapped reads
outdir: A tarball of the Snippy output directory for input into Snippy-core if required
Question
We could go through all of the variants like this and read them out of a text table, but this is onerous and doesn’t really give the context of the changes very well.
In order to have a visualisation of the SNPs and the other relevant data, in Galaxy we can use JBrowse
JBrowse Tool with the following parameters
Select the reference genome: wildtype.fna
This sequence will be the reference against which annotations are displayed
We will now set up three different tracks - these are datasets displayed underneath the reference sequence
(which is displayed as nucleotides in FASTA format). We will choose to display the sequence reads (the .bam file)
the variants found by snippy (the .gff file) and the annotated reference genome (the wildtype.gff)
SnippySnippyminus button to see sequence reads and their coverage (the grey graph)plus button to see
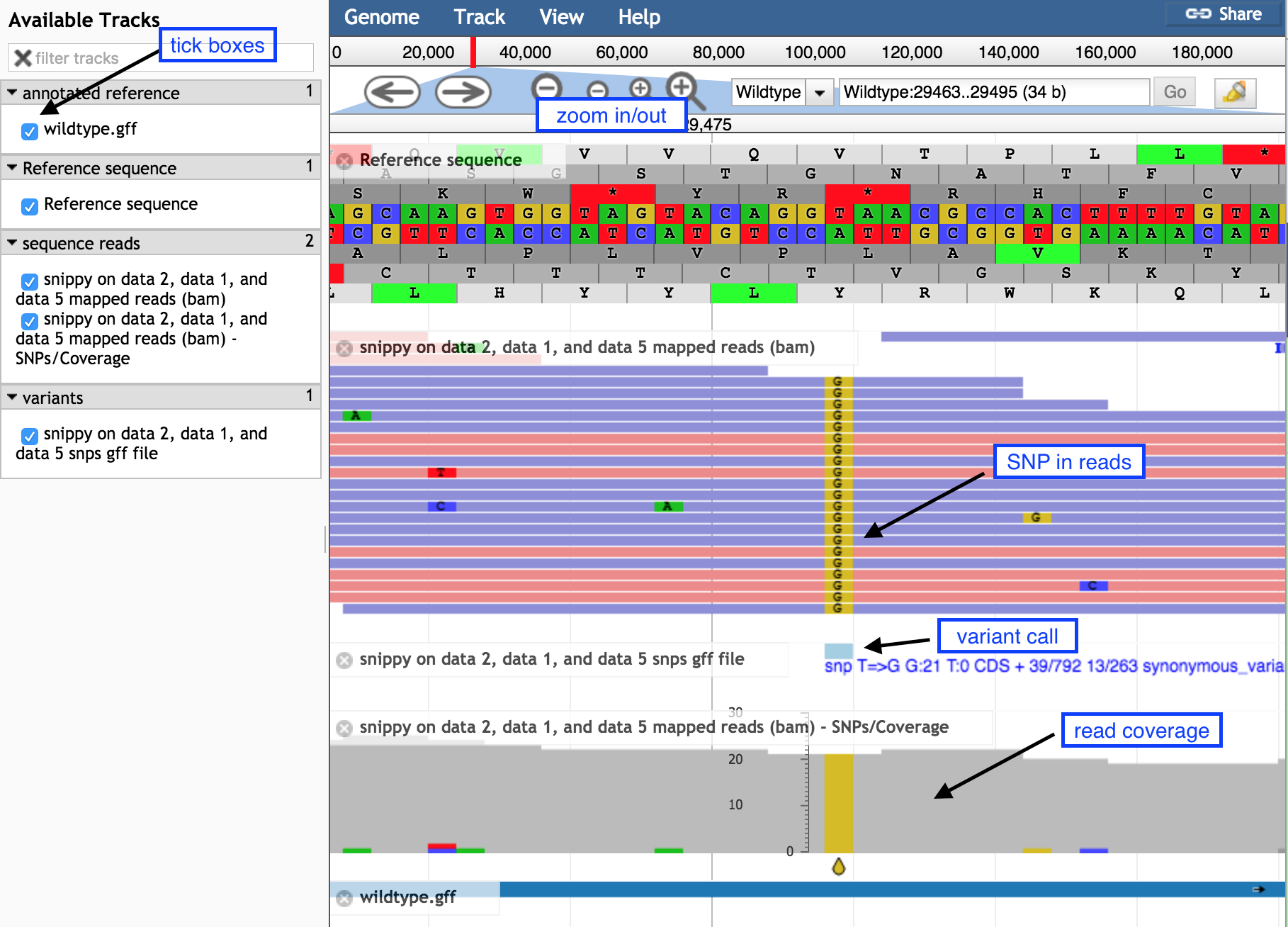
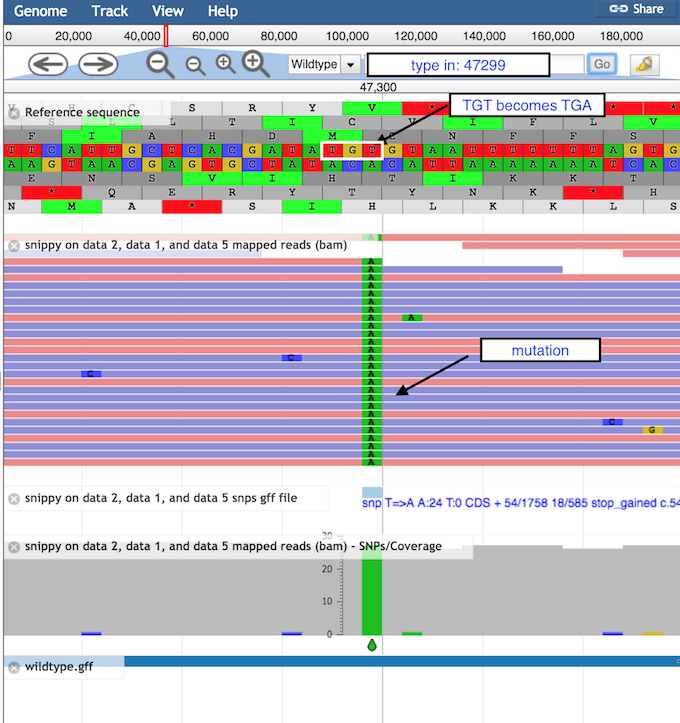
Question
By running a tool such as Snippy on your read files and reference genome,
we can find where the biologically important changes between genomes
of different strains occur and perhaps what they mean to the phenotype
We used a tool called Snippy to call variants between our reads and our reference genome.
As our reference genome had annotations, we could see what effect the changes have on the features as annotated in the reference and therefore make inferences on the possible changes to the phenotype.
We used the JBrowse genome browser to visualise what these changes look like